News
-
 31 March 2026
31 March 2026
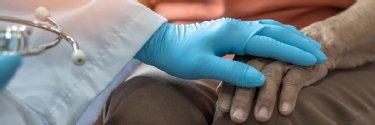
Aussie AI health-tech Heidi aims to cure clinical burnout
-
 31 March 2026
31 March 2026
Delta in-flight connectivity takes off with Amazon Leo
-
 31 March 2026
31 March 2026
High Court dismisses judicial review against eVisa system
-
 31 March 2026
31 March 2026
CityFibre launches 8.5Gb service across wholesale multi-gig network
News Archive
-
March 31, 2026
31
Mar'26
Aussie AI health-tech Heidi aims to cure clinical burnout
Armed with over $100m in funding, Melbourne-based Heidi is building its own AI models and launching wearable hardware to automate documentation and reduce the administrative burden on doctors
-
March 31, 2026
31
Mar'26
Delta in-flight connectivity takes off with Amazon Leo
Global airline looks to AWS satellite division to introduce connectivity on hundreds of aircraft, starting with an initial installation on 500 aircraft in 2028, working to expand its Wi‑Fi and seatback experiences
-
March 31, 2026
31
Mar'26
High Court dismisses judicial review against eVisa system
The High Court rules that the Home Office is acting lawfully in refusing to issue alterative proof of immigration status outside of its electronic visa system, but both the judge and the department accepts that those affected by data quality and ...
-
March 31, 2026
31
Mar'26
CityFibre launches 8.5Gb service across wholesale multi-gig network
UK’s largest independent full-fibre platform provider makes the next step in its roll-out of its 10Gb-capable network, and claims strong customer growth continuing as it approaches one million users
-
March 31, 2026
31
Mar'26
Cato Networks unveils modular adoption model for SASE platform
SD-WAN, SSE, AI security and UZTNA standalone modules at heart of a converged platform that unlocks operational simplicity, resilient connectivity and stronger security
-
March 31, 2026
31
Mar'26
CMA to launch strategic market status investigation into Microsoft; Amazon Web Services off the hook
CMA to investigate whether Microsoft should be given strategic market status. Amazon escaped, but both companies will need to make changes to egress fees and interoperability
-
March 31, 2026
31
Mar'26
Virgin Media O2 accelerates UK 5G upgrade programme
UK operator inks multi-year agreements with strategic essential equipment partners to upgrade radio access network to next-generation 5G technology, boosting capacity, coverage and reliability
-
March 31, 2026
31
Mar'26
Shrinking PQC timeline highlights immediate risk to data security
Google’s decision to move up its timeline for migration to post-quantum cryptography highlights that some of the cyber security risks posed by quantum computing are already reality
-
March 31, 2026
31
Mar'26
Agoda scales AI strategy, opens new APAC tech hub
The digital travel platform has set its sights on becoming an AI-powered travel companion as it changes how it builds software and moves its tech workforce into a new facility in Bangkok
-
March 30, 2026
30
Mar'26
Orange steams ahead in French railway connectivity
Analysis of data across 20 major French rail corridors reveals sharp operator disparities in throughput, latency and quality of experience
-
March 30, 2026
30
Mar'26
Cambridge Mobile Telematics lands $350m strategic investment
Major investment in automotive telematics company, accompanied by new long-term commercial agreements, seeks to expand global road safety platform and real-time AI risk models
-
March 30, 2026
30
Mar'26
Mobile network operators urged to help commercialise 5G live production
Private 5G connectivity provider gains support from broadcasters and tech giants to calls on mobile network operators to enable on-demand network quality for live 5G broadcasts
-
March 30, 2026
30
Mar'26
Interview: Thierry Martin, head of enterprise data and analytics, Toyota Motor Europe
A sketch artist by night, and a vehicle engineer by training, Toyota Europe’s data chief is bringing elements of both capabilities to bear in delivering better data insights and building a foundation for AI
-
March 30, 2026
30
Mar'26
Stop Scams steps up to online fraud challenge
After years of putting the building blocks in place, Stop Scams is ready and able to react quickly to fight emerging fraud threats
-
March 30, 2026
30
Mar'26
Starlink reshapes satellite communications as industry enters terabit era
Lower-cost capacity, rapid scaling and improved service quality are all factors resetting expectations across the satcom market, with the entire ecosystem being pushed to innovate, differentiate and rethink strategic positioning
-
March 29, 2026
29
Mar'26
Advancing to the next frontier of AI
As AI agents move faster than software made for human users, both digital tooling and silicon architecture need to be redesigned to reduce latency and power bottlenecks, according to chief scientists of Nvidia and Google
-
March 27, 2026
27
Mar'26
Pave Space raises $40M to hasten satellite deployment
Swiss startup building orbital transfer vehicles capable of delivering satellites to operational orbits in under 24 hours, addressing a growing bottleneck in the space economy
-
March 27, 2026
27
Mar'26
UK government lacks ambition to fight tax fraud, says PAC
The Public Accounts Committee says the UK government has dropped the ball on the use of data analytics to tackle tax fraud and error, as the public purse haemorrhages billions of pounds
-
March 27, 2026
27
Mar'26
Lloyds admits coding fault exposed customer transactions
The bank has responded to the Treasury Committee’s request for information on a major data breach in its banking app
-
March 27, 2026
27
Mar'26
Virgin Media Business Wholesale accelerates high-capacity delivery in the UK
Fixed wholesale connectivity arm of leading UK operator announces biggest ever upgrade to its wholesale network, delivering 32% Ethernet lead time reduction and 10G delivery accelerated by up to 40 days
-
March 27, 2026
27
Mar'26
Flaws in government procurement show in HMRC £473m AWS award
After a rushed contract award with only one bidder and a tender notice ‘for hyperscalers only’, critics call for live oversight on government contracts, claiming the procurement is unfair and likely to be expensive
-
March 27, 2026
27
Mar'26
Antevia joins O-RAN Alliance to simplify enterprise 5G deployment
Private network technology provider looks to strengthen commitment to development of open, interoperable and simplified enterprise cellular deployments and scale enterprise 5G
-
March 27, 2026
27
Mar'26
Second Post Office Capture conviction referred to appeal court
Conviction of 30 years has been referred to the Court of Appeal by the Criminal Cases Review Commission
-
March 27, 2026
27
Mar'26
Colt announces subsea, terrestrial network routes
Digital infrastructure company reveals plans to launch international connectivity routes connecting the US West Coast to Asia, marking the latest phase of its major global network expansion
-
March 27, 2026
27
Mar'26
Capita left to deal with 13,000 civil service pension cases over a year old
More details of the Civil Services Pension Scheme administration backlog left to Capita revealed in parliamentary committee hearing
-
March 27, 2026
27
Mar'26
EU Parliament rejects Chat Control message scanning
MEPs vote down proposals to allow US tech companies to continue scanning private messages for illegal content
-
March 26, 2026
26
Mar'26
Oracle opens Sydney customer excellence centre to boost AI adoption
Facility will help organisations across Australia and Oceania navigate technology challenges and turn AI experimentation into business value
-
March 26, 2026
26
Mar'26
Oracle Cloud Infrastructure: The bare metal facts
The Oracle Cloud Infrastructure appears to have more in common with datacentre hosting than with public infrastructure-as-a-service providers
-
March 26, 2026
26
Mar'26
Nokia joins Linx as technical partner for London network refresh
Internet exchange based in UK capital completes project refreshing its 17-site interconnected network in London, with global comms tech provider selected as the technical partner to support development
-
March 26, 2026
26
Mar'26
Proptivity, Telehouse team for reliable indoor 4G, 5G in London workplaces
Datacentre service provider and indoor mobile infrastructure firm look to address connectivity blind spot inside modern office buildings, enabling scalable indoor 4G, 5G across London workplaces via neutral host model
-
March 26, 2026
26
Mar'26
Post Office still can’t find evidence for 1,400 scandal redress claimants, while people die waiting
Finding evidence for events that took place decades ago is a challenge for many subpostmasters seeking compensation through the Horizon Shortfall Scheme
-
March 26, 2026
26
Mar'26
Filtronic unveils DIFI service to enable virtualised satellite ground stations
Space, defence and telecoms infrastructure provider intros high-bandwidth digital intermediate frequency tech designed to support the transition to fully virtualised satellite ground stations
-
March 26, 2026
26
Mar'26
UAE positions cyber security as pillar of national resilience and digital growth
Strategic investment and coordination reinforce the country’s ability to withstand complex cyber threats
-
March 26, 2026
26
Mar'26
Bank of England IT project offers lessons for wider government
The UK central bank’s core IT system replacement project surprised MPs, who were unaccustomed to reviewing success stories – its achievement could potentially serve as a model for future government IT initiatives
-
March 26, 2026
26
Mar'26
Connectivity to the fore as Sunderland commits to 2035 digital strategy
North eastern English city expands its smart city commitments with 2035 strategy to ensure every resident can “thrive in an increasingly digital world”
-
March 26, 2026
26
Mar'26
Agentic bots and synthetic identities fuel surge in fraud
LexisNexis Risk Solutions warns of a 450% rise in agentic traffic and an eight-fold increase in synthetic identity fraud as cyber criminals scale automation to bypass security controls
-
March 25, 2026
25
Mar'26
US government launches Bureau of Emerging Threats
The US’ Bureau of Emerging Threats sits within the State Department and will supposedly help address national security threats arising from cyber attacks, the weaponisation of space and other emerging technologies
-
March 25, 2026
25
Mar'26
Google targets 2029 for post-quantum cyber readiness
Google sets out a timeline for its migration to post-quantum cryptography, saying it will complete its migration before the end of the 2020s
-
March 25, 2026
25
Mar'26
Amazon Web Services bags Fujitsu’s HMRC loss
US tech giant wins contract to run three datacentres for the government department after cutting ties with Fujitsu
-
March 25, 2026
25
Mar'26
SES, K2 Space further meoSphere satellite network
Next-generation medium Earth orbit multi-mission constellation targeted at meeting fast-growing commercial and defence needs with flexible scaling approach
-
March 25, 2026
25
Mar'26
Emergency Microsoft, Oracle patches point to wider cyber issues
Emergency out-of-band patches from Microsoft and Oracle signal underlying security issues around update cycles and patching, and identity security and zero-trust, says the community
-
March 25, 2026
25
Mar'26
Oracle applications chief sees enterprise AI agents as task-specific helpers
At Oracle AI Summit in London, Steve Miranda, executive vice-president of Oracle applications development, discussed Oracle’s Fusion Agentic Applications, including how should AI agents be used, how is it priced and what safeguards are being put in ...
-
March 25, 2026
25
Mar'26
Google Cloud, Openreach expand connectivity collaboration
UK’s largest broadband operator implements AI to accelerate high-speed internet access and cut carbon, in an expanded collaboration to accelerate Openreach’s sustainability and connectivity goals
-
March 25, 2026
25
Mar'26
Akamai launches AI Grid intelligent orchestration
Cyber security and cloud computing company unveils global-scale implementation of AI Grid, intelligently routing artificial intelligence workloads across edge, regional and core footprint to balance latency, cost and performance
-
March 25, 2026
25
Mar'26
UK government boosts digital access for more than a million people
Since its launch last year, the UK government’s Digital Inclusion Action Plan has consistently met its targets, helping over one million people access the digital world
-
March 25, 2026
25
Mar'26
Why AI agents are one prompt away from ransomware
As AI adoption advances beyond chatbots, security leaders are up against rogue AI agents mirroring threat actors and a generational skills gap as security operations teams become overly dependent on AI
-
March 25, 2026
25
Mar'26
Designing water‐smart AI datacentres in GCC and MENA
As Gulf countries race to build AI capacity in one of the world’s most water‐stressed regions, policymakers and operators are rethinking how they power and cool datacentres without exhausting critical water resources
-
March 25, 2026
25
Mar'26
Ciena upgrades subsea cable throughput for Meta, Lightstorm
Optical connectivity provider’s technology is used to set transmission record for across a transpacific submarine cable, enabling next-generation cloud and AI connectivity between Japan and Australia
-
March 25, 2026
25
Mar'26
The UAE CIO: From technology operator to digital value architect
As AI, sovereign cloud and regulation reshape the Gulf’s technology landscape, CIOs in the UAE are being pushed beyond infrastructure management into a role that blends strategy, governance and digital trust, says CIO and technology expert Umesh ...
-
March 25, 2026
25
Mar'26
Cambridge MC boosts comms procurement with Carrier Club acquisition
International consulting firm swoops for ’unique’ procurement-as-a-service organisation to offer greater value to its clients in challenging times for businesses


